This web page was produced as an assignment for Bioinformatics 490-006, an undergraduate and graduate student course at Southern Illinois University Edwardsville.

BIOINFORMATICS
Resources and Outside Information
Protein Domains
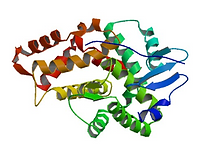
1DXX
N-Terminal Actin Binding Domain.
The N-Terminus of the Dystrophin protein bind to F-actin, while the C-terminus binds to the dystrophin-associated glycoprotein (DAG) complex.
Dystrophin links the actin-base cytoskeleton of muscle cells to the extracellular matrix.
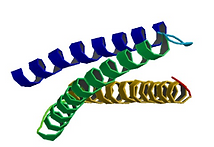
1EG4
WW Domain protein structure in complex with a beta-dystroglycan peptide
The C-terminus of the dystrophin protein binds to beta-dystroglycan.
The WW domain is adjacent with two EF-hand-like domains which associate with beta-dystroglycan
1EG3
WW Domain protein structure in complex with a beta-dystroglycan peptide
Dystrophin protein and beta-dystroglycan are components of multimolecular complex dystrophin-associated gycoprotein complex, which links muscle cells cytoskeleton to the extracellular lamina.
This figure shows another WW domain of dystrophin, which couples the proteins using a proline-rich binding site in beta-dystrogylcan.
3UUN
N-Terminal first spectrin repeat
The dystrophin protein works to link the cyctoskeleton to the extracellular matrix by having its N-terminal actin binding domain, and C-terminal protein binding motifs. Between is several spectrin-repeat domains.
This figure shows the first spectrin repeat domain of dystrophin. The structure contains a 3-helical fold, along with and extention of the spectrin repeat domain, which is show to stabilize the protein.
Norwood, F. L., Sutherland-Smith, A. J., Keep, N. H., & Kendrick-Jones, J. (2000). The structure of the N-terminal actin-binding domain of human dystrophin and how mutations in this domain may cause Duchenne or Becker muscular dystrophy. Structure, 8(5), 481–491. doi:10.1016/s0969-2126(00)00132-5
Huang, X., Poy, F., Zhang, R., Joachimiak, A., Sudol, M., & Eck, M. J. (2000). Nature Structural Biology, 7(8), 634–638. doi:10.1038/77923
Huang, X., Poy, F., Zhang, R., Joachimiak, A., Sudol, M., & Eck, M. J. (2000). Nature Structural Biology, 7(8), 634–638. doi:10.1038/77923
Muthu, M., Richardson, K. A., & Sutherland-Smith, A. J. (2012). The Crystal Structures of Dystrophin and Utrophin Spectrin Repeats: Implications for Domain Boundaries. PLoS ONE, 7(7), e40066. doi:10.1371/journal.pone.0040066

Wild-Type

MDX
RNA-seq on Regenrated mdx Mouse Muscle Tissue
The mdx mouse is often use to research Duchenne Muscular Dystrophy. These mice have a point mutation in the DMD gene, causing an amino acid change inducing a nonsense mutation. This mutation produces non-functional dystrophin proteins.
Gene expression was analysed in 16-week regenated mdx mice to discover any pathway made to save muscle integrity in absence of the DMD protein.
The DMD gene is highlighted orange in the volcano plots. In the Wild-Type analysis, DMD is highly expressed, while in the MDX plot, it has low expression.
Sequence Alignment
The DMD gene spans millions of base pairs in Homo sapians and many of its homologs in other species. The 30 reference sequence listed on NCBI were aligned in UGene using muscle and further analysed. Sequences included variants with mutations that induce Becker Muscular Dystrophy, and predicted sequences using Gnomon.
Below is just a snip of one of the most conserved parts of the 14,577 base pair alignment

Protein Alignment Across Species
Another alignment was made of conserved DMD proteins from different species. Canis lupus protein sequence was most similar to Homo sapiens, with a 95% similarity.

DMD Pairwise Alignment: Human and Chimp
A pairwise alignment between Homo sapiens and Pan troglodytes was also made using NCBI Blast, and found that the partial DMD sequences were over 99% similar with 0 gaps.

Stephen F. Altschul, Thomas L. Madden, Alejandro A. Schäffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of protein database search programs", Nucleic Acids Res. 25:3389-3402
Using Microarray analysis to detect DMD mutations
Current methods of mutation detection can be time consuming and inaccurate, largely due to the size of the DMD gene. Comparative Genomic Hybridization (CGH) microarrays can be used to detect mutations and screen for DMD and BMD and can be used in clinical molecular testing laboratories.
In this study, a retrospective analysis of 29 patients were done on 15 males and 14 females of varying mutations. Multiplex PCR and Southern blotting were done blinding to characterize the mutation types, before microarray analysis could be compared.
Not only was it found that microarray analysis was extremely accurate at detecting mutations, it also has a quick turnaround of only 7-10 days, and is cheap and readily available
Hegde, M. R., Chin, E. L. H., Mulle, J. G., Okou, D. T., Warren, S. T., & Zwick, M. E. (2008). Microarray-based mutation detection in thedystrophingene. Human Mutation, 29(9), 1091–1099. doi:10.1002/humu.20831

The figure above shows a sample of what a CGH microarray result looks like. A 240-bp sample of the DMD gene was taken to show CGH probe distribution, with each thin blue line representing one probe. the Refeq gene at the bottom is the reference for Exon 44.

This figure show a sample clinical CGH microarray. Exons 1 through 79 were mapped for 8 different samples of DMD mutations.
1: Male deletion of exon 18
2: Male with deletions of exon 45 - 54
3: Male with duplication of exon 2-4
4: Male with duplication of exons 35 -4
5: Male with 33kb deletion of intron 1 and an 11kb deletion of intron 2
6: Female with deletion of exons 49-50
7: Female with duplication of exon 44
8: Female with duplication of exon 8-11