This web page was produced as an assignment for Bioinformatics 490-006, an undergraduate and graduate student course at Southern Illinois University Edwardsville.
PUBLICATIONS
Recent Publications on DMD
Identifying DMD mutations using multiple ligation probe amplification (MLPA)
New ways to screen for genetic disorders are always being studied to keep cost down and health care accessible and moving forward. In this study multiplex PCR and MLPA analysis were compare to determine their accuracy at detecting specific mutations in the DMD gene.
It was found that MLPA analysis was a simple, accurate, and rapid method in detecting DMD mutations. Although some inaccuracies were found in the case of a known female carrier in which no mutation was detected.
Gatta, V., Scarciolla, O., Gaspari, A. R., Palka, C., De Angelis, M. V., Di Muzio, A., … Stuppia, L. (2005). Identification of deletions and duplications of the DMD gene in affected males and carrier females by multiple ligation probe amplification (MLPA). Human Genetics, 117(1), 92–98. doi:10.1007/s00439-005-1270-7

Fig. 1 Shows a side by side comparison of the analysis of 6 patients using PCR and MLPA. This shows that MLPA can give you a rapid overview of genetic mutations in a patient.
Editing Genes of Dystrophic Mice Muscle Cells and Muscle Stem Cells In Vivo
Removing exons in a frame shifted mutated DMD gene can restore partial function of the DMD protein. Adeno-Associate Virus (AAV) of Clustered Regularly Interspaced Palindromic Repeats (CRISPR)-Cas9 were used with a guide RNA on a mutated DMD gene on exon 23 to restore reading frames.
It was found that AAV-Dmd-CRISPR-treatment partially restored muscle functions in mdx mouse muscle.
Tabebordbar, M., Zhu, K., Cheng, J. K. W., Chew, W. L., Widrick, J. J., Yan, W. X., … Wagers, A. J. (2015). In vivo gene editing in dystrophic mouse muscle and muscle stem cells. Science, 351(6271), 407–411. doi:10.1126/science.aad5177
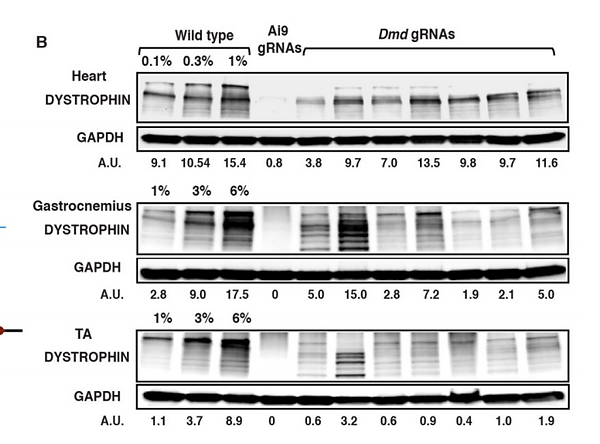
Fig. 2 shows a western plot detecting dystrophin protein expression in wild type mice, mice treated with an AI9 guide RNA and mice treated with Dmd guide RNAs. GAPDH was used as a signal intensity standard, determined by densitometry.
It is shown that AI9 guide RNAs do little to increase expression the dystrophin, while Dmd guide RNAs can partially restore expression of the protein.
Normal and altered pre-mRNA processing
Dmd is a large gene that required accurate mRNA processing to produce function dystrophin proteins. This article overviews mechanisms involve in Dmd splicing, and proposes how alternative splicing can modify disease in patients by changing the outcome of the primary effect.
All 79 exons of Dmd are expressed in adult skeletal tissue, however splicing makes it possible for a number of protein variants to arise in different tissue and during different developmental stages of life.
Studying splicing of Dmd can provide insight on diseases and dystrophins cellular processes. More most be done to fully understand how each factor plays a role in RNA processing.
Tuffery-Giraud, S., Miro, J., Koenig, M., & Claustres, M. (2017). Normal and altered pre-mRNA processing in the DMD gene. Human Genetics, 136(9), 1155–1172. doi:10.1007/s00439-017-1820-9

Fig. 3 shows different patterns of alternative splicing. Strengths of 5' and 3' splice sights (ss) of Dmds 79 exons were calculated with the bioinformatics tool Human Splicing Finding (HSF). These scores are represented on the 5' and 3' wheels.
Pictures b through g in figure show methods of splicing.